Even though protein based biosensors have been successfully developed, they have typically used naturally occurring enzymes that have not evolved for the purpose of being used in a biosensor. Similarly, the hunt for naturally occurring bioelectronic components does not reveal structures optimised for assembly and deployment in circuitry. A powerful tool currently being used to expand the family of biomaterials and integrate them with transduction elements is protein engineering.
Starting from the basic concept that a biosensor for in vivo diagnostics requires an analyte biorecognition element, for example an enzyme, antibody or other binding protein, biosensor design begins with that element. The vision converges towards modular molecular-engineering in which integrated signal transduction function and binding site characteristics are engineered in tandem with self-assembling nanostructures or modular self-immobilising peptide sequences. These design principles have allowed protein isolation without many of the normal downstream steps and avoiding chemical modification, while achieving stabilisation of the proteins without such stringent cold chain needs.
Protein engineering allows both structure and function to be embedded into a fusion protein and optimised to become a novel bioanalytical component. For example, a construct of an enzyme or antibody with honeybee silk protein appended with fluorescent protein can be expressed in E. coli and spun to fibres which can be fashioned into swabs and included in a design for a self-indicating in vitro diagnostic.
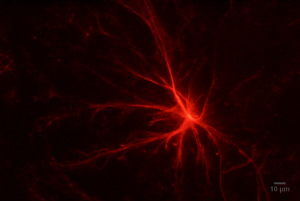
Monomeric Red Fluorescent Protein (mcRFP)-Q77 silk fusion protein fibre formation
Using synthetic biology we have also used properties of energy transfer to design a self referencing firefly luciferase enzyme which measures ATP and taking into account the distance between the redox site and the electrode have engineered redox proteins to allow direct electron transfer between protein and electrode. Hybrid protein-redox polymer structures have been achieved whose oxidation state and charge storage can be controlled by electrode polarisation voltage.